Single-cell spatial transcriptomic analysis of human skin anatomy
Key Points:
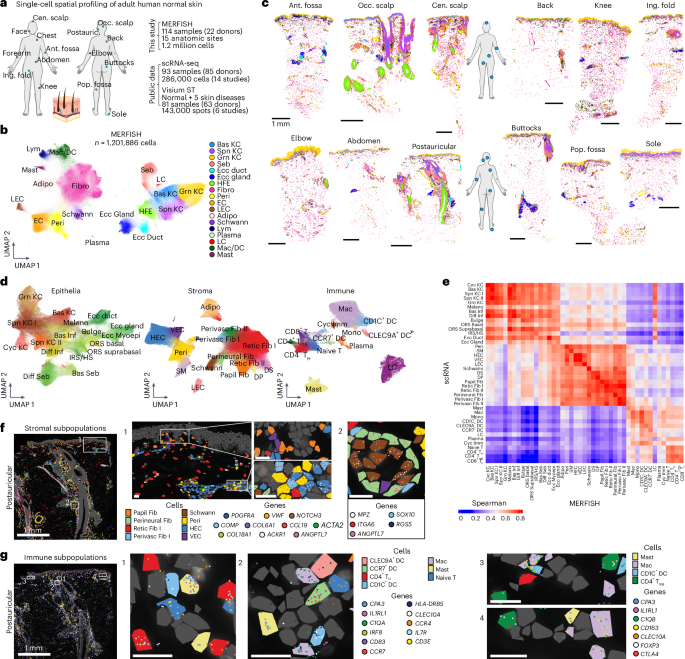
- Researchers generated a comprehensive single-cell spatial transcriptomic atlas of normal human skin using MERFISH on 114 samples from 22 donors across 15 body sites, identifying 45 distinct cell populations including epithelial, stromal, and immune subtypes with precise spatial localization.
- Analysis revealed stereotypic cellular distribution patterns and ten multicellular neighborhoods that define skin microanatomy, with site-specific variations in cell diversity and density, notably higher diversity in hair-dense and flexural regions and unique features in the sole.
- The study uncovered extensive ligand-receptor interactions within neighborhoods, particularly in the immune-enriched perivascular (PERIVASC I) neighborhood, where TNF signaling was shown to maintain CCL19+ perivascular fibroblast identity, validated by functional assays in human skin explants.
- Integration with public spatial transcriptomic data demonstrated that skin diseases (e.g., atopic dermatitis, psoriasis, hidradenitis suppurativa, basal and squamous cell carcinomas) are associated with expansion and transcriptional remodeling of homeostatic neighborhoods, especially the PERIVASC I neighborhood, indicating its role as an immunomodulatory spatial domain.
- The atlas and associated computational tools provide a valuable resource for understanding human skin biology, spatial cell-cell communication, and disease-associated microanatomical remodeling, with implications for targeted therapeutic strategies.